Dendrites are the tree-like arborizations of neurons through which they receive input from other neurons. This compartmentalized anatomy has given rise to the idea that individual dendrites process information independently from each other and from the soma. This idea has driven many studies on dendrite function during the last 30 years, mostly in brain slices. But do neurons and dendrites make use of these compartmentalized computations in the intact, living brain?
Recent advances in microscopy and voltage imaging have made it possible to address this question more directly than ever before. Specifially, there has been a recent streak of technically impressive papers on dendritic imaging in the hippocampus of awake mice. In these papers from the labs of Adam Cohen, Attila Losonczy, and Balázs Rózsa, hippocampal pyramidal neurons together with their dendrites are imaged in the behaving mouse, using both calcium and voltage imaging (Gonzalez et al., 2026; Lee et al., 2026; Moore et al., 2025; Wu et al., 2026). Let’s have a closer look!
Paper 1: Two-photon random-access voltage and calcium imaging of CA1 dendrites
Movement-stabilized three-dimensional optical recordings of membrane potential and calcium by Gonzalez, Terada, et al., Neuron, 2026.
Online motion correction for fast dendritic two-photon imaging
Recording both voltage and calcium in 3D is a “holy grail” for dendritic physiology. However, this requires scanning only the dendrites without their surrounding at high speed and repetition rate, making the recording susceptible to artifacts due to brain motion that make such data unusable. To fix this problem, the laser scan pattern needs to be corrected online with millisecond precision to compensate for brain motion. Here, the Rózsa and Losonczy labs developed a new approach that they termed 3D real-time motion correction (3D-RTMC).
Real-time motion correction is highly challenging, and a bright and stable object needs to be used as reference. Earlier work from the lab of Angus Silver injected bright beads for that (Griffiths et al., 2020). Here, the authors took advantage of the single-cell electroporation experts in the Losonczy lab (Gonzalez et al., 2025) and electroporated a cell close to the cell of interest with a fluorescence-expressing plasmid in order have a closeby bright object as a reference for online motion correction. This method allowed them to track the same dendritic branches in a head-fixed mouse running on a treadmill with sub-micron precision using simultaneous voltage and calcium imaging.
Decoupling of calcium vs. voltage with higher dendritic branch order
The authors found that as signals traveled further from the soma, voltage and calcium dynamics became more and more distinct. Proximal dendrite activity mirrored somatic activity, but distal branches showed a progressive decoupling, with branch order, not just distance, determining how well calcium reflected the underlying voltage. A limitation here is that they used the red-shifted jRGECO1a as a calcium indicator (which was necessary so it could be combined with the green voltage sensor ASAP6). jRGECO1a reports calcium changes in a nonlinear way; this nonlinearity is likely to result in missing small dendritic events (Rupprecht et al., 2025).
Local dendritic voltage signals
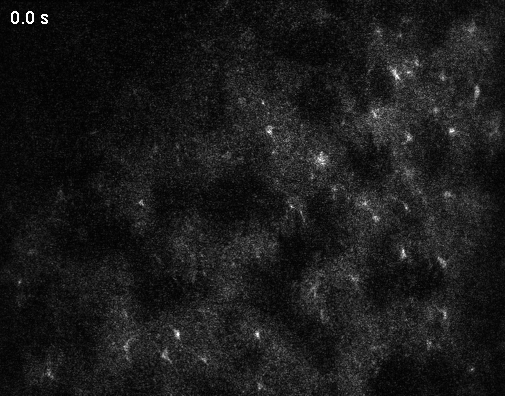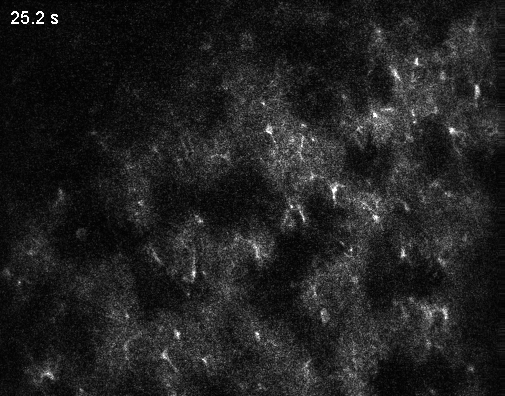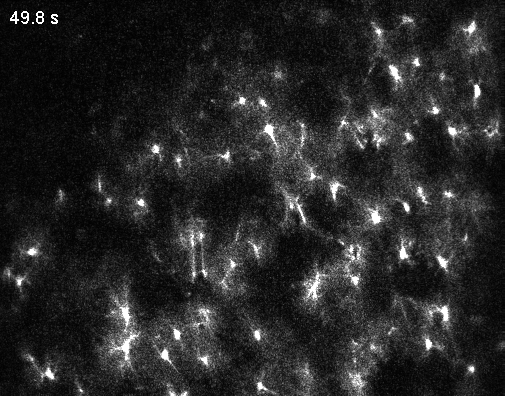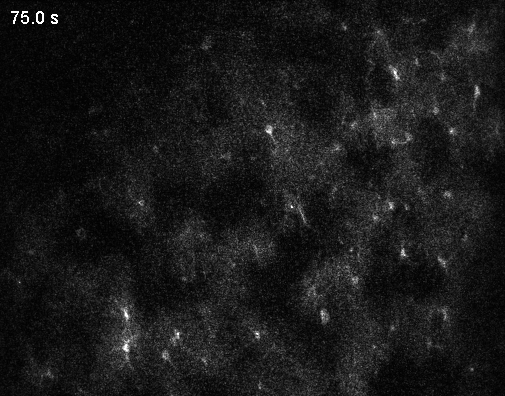
Interestingly, the authors describe an abundance of local calcium signals in dendrites independent of somatic activity (Figure 2C,K,L; Supplementary Figure 4). However, this important observation is only briefly described and not discussed in depth. For example, Figure 2L shows that dendrites and soma are very likely to be coactive; but if the dendrite is active, the soma is recruited (“s|d”) with a higher probability than the other way around (“d|s”). This is the opposite of what is reported in the paper by (Lee et al., 2026) discussed below and would be worth discussing. – For clarity, the Lee et al. paper came out after the Gonzalez et al. manuscript was published, so the authors had no way to discuss this finding.
Paper 2: Prism-based one-photon voltage imaging of CA1 dendrites
Fast dendritic excitations primarily mediate back-propagation in CA1, by Lee, Park, et al., bioRxiv, 2026.
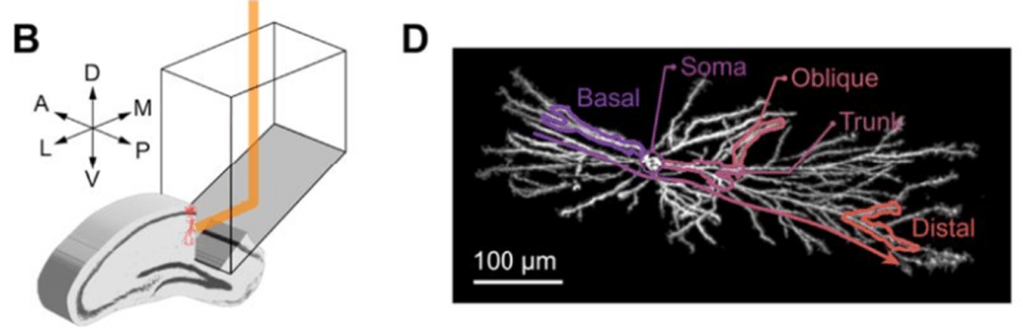
Prism-based voltage imaging of hippocampal dendrites using one-photon microscopy. Excerpt of Figure 1 from the preprint by Lee et al., under CC BY-NC-ND 4.0 license.
One-photon voltage imaging of individual CA1 neurons
The Cohen lab adapted a previously published design (Redman et al., 2022), using a side-on microprism and kHz-rate voltage imaging with DMD-based structured illumination to record from a large fraction of the entire dendritic tree of CA1 neurons (Lee et al., 2026). This manuscript provides to the best of my knowledge the most comprehensive functional voltage recording of a single neuron in a behaving animal to date.
I’m very impressed by how they combined high-end optics, genetic tools, and very complex data analysis for these experiments in behaving head-fixed mice. From a scientific perspective, the paper is very difficult to summarize because it covers several major questions of neurophysiology in an almost comprehensive manner. And I believe that the choice of the paper title could be improved because it highlights only one aspect of their work.
A note on data analysis
Sometimes, there are Methods sections that are instructive by being rich in interesting details and full of complex processing steps which correct for confounds that you would rarely think of in the beginning: this is one of them. One detail that I found a bit confusing is that they regressed out brain motion (derived from the motion correction algorithm) from the dendritic voltage signals. This is desribed at the bottom of the “Voltage signal extraction” section in Methods. Would this not artificially subtract any dendritic signals related to the movement of the animal? I am a bit skeptic about this methodological detail.
Anti-correlation of apical vs. basal dendrite activity
Lee et al. find that subthreshold voltage is primarily explained by global activity across the entire arbor (~55% of variance), followed by a pattern that they term “see-saw”, where apical and basal dendrites are anticorrelated (~20% of the variance; clearly visible in the movie below). I really like how they collect evidence that this see-saw pattern is most likely determined by laminar-specific inputs in CA1. They do so by comparing the spatial extent of spontaneous activity patterns (large extent) with spatial patterns generated by local dendritic optogenetic depolarization (smaller extent).
Supplementary Movie 3 from the preprint by Lee et al., under CC BY-NC-ND 4.0 license.
Evidence against prominent local voltage events in dendrites
Interestingly, Lee et al. do not find evidence for prominent local voltage events in dendritic compartments, in contrast to the data from the Losonczy/Rósza labs (Gonzalez et al., 2026); and only limited evidence for dendrite-specific place fields, in contrast to a previous calcium imaging study (Moore et al., 2025).
Instead, they find that most “spikes” seen in dendrites originate from back-propagating action potentials (bAPs) in the soma (90%). With this, they identify the soma and not the apical dendrite as primary site of origin for bAPs and complex spikes. In the end, of course, all somatic spikes are caused by the cumulative integration of dendritic inputs. However, this study suggests that the amplification of those signals is primarily a consequence of successful somatic firing and not of amplification in the dendrites, contrasting with existing theories focusing on dendrite-localized amplification (Larkum, 2013; Major et al., 2013; Poirazi and Papoutsi, 2020).
Backpropagation enhanced by local depolarization
The authors also find that apical depolarization enhances bAP propagation into the apical dendrite, which would make bAP propagation specific to sites of prior activity. This finding is not very surprising and confirms earlier work in slices; but it strengthens the idea of bAPs as a learning signal for local dendritic plasticity and is a very important piece of evidence.
Many more biophysical and physiological findings are in the paper, and every paragraph is interesting to read. Highly recommended!
Paper 3: Simultaneous calcium and voltage imaging across the dendritic tree
A dendrite-resolved, in vivo transfer function from spike patterns to Ca2+, by Wu, Lee, et al., bioRxiv, 2026.
One-photon imaging of voltage and calcium in individual neurons
This sister paper from Adam Cohen’s lab also uses prism-based one-photon imaging, recording not only voltage but also calcium events of hippocampal pyramidal cells across a large fraction of the dendritic tree (Wu et al., 2026). To this end, they use a slightly different optical setup, with a spinning disk-based design. They focus on CA2 pyramidal neurons, not primarily for scientific reasons but because expression is much stronger in CA2 for most AAV serotypes (Alexander et al., 2024). Imaging calcium and voltage simultaneously is one of the most interesting experiments for researchers like myself who worked on interpreting calcium imaging data for a long time (Rupprecht et al., 2021, Rupprecht et al., 2025). Achieving such a dual recording across a large fraction of the dendritic tree definitely exceeded my expectations.
A biophysical rule to convert voltage to calcium
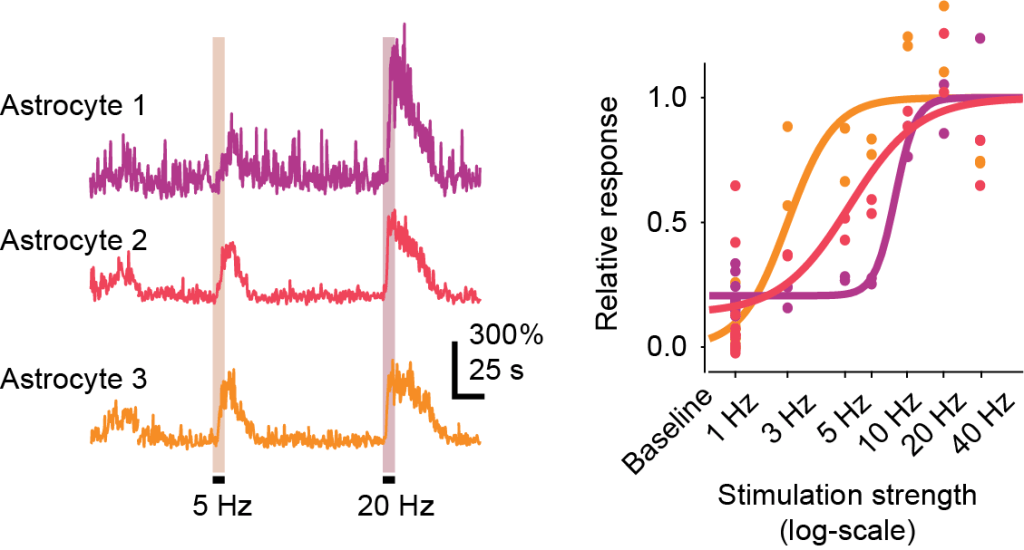
Wu et al. find that the mapping from voltage to calcium is remarkably predictable and follows a hierarchical activation pattern. To explain the data, they fit a simple sigmoidal model to the voltage-calcium transfer function. They find that the inflection point of this transfer function (which they term V0 in their paper) becomes larger for smaller dendritic and especially apical compartments. They argue that this might be a mechanism to avoid noisy dendritic calcium signals due to local depolarizations from synapses. The combination of acquiring such a complex dataset and modeling it in such simple and straightforward terms is quite beautiful.
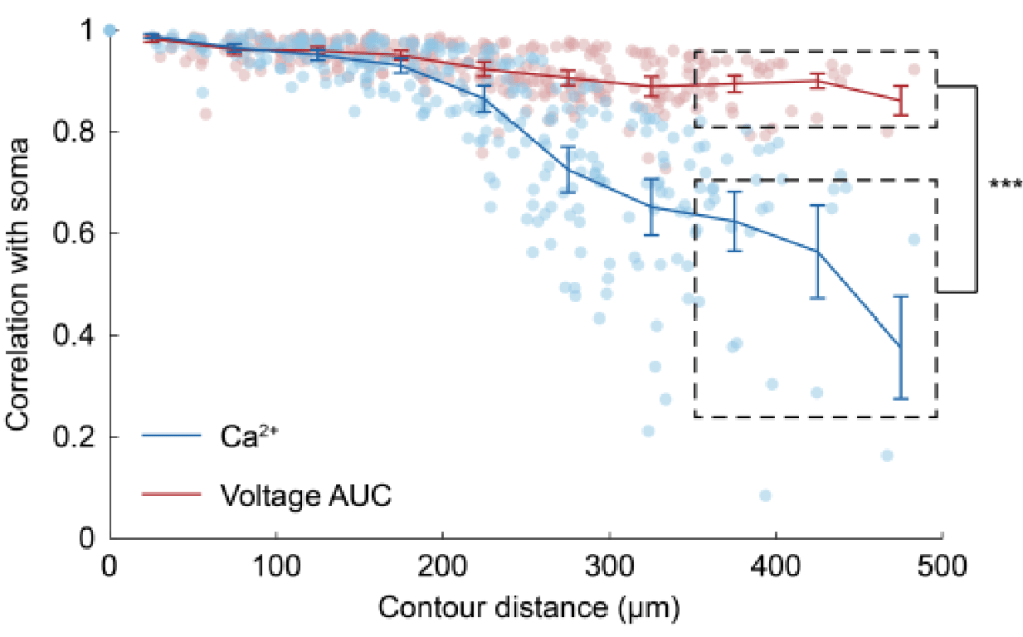
Similarity to somatic activity decreases with distance from soma for calcium signals and less so for voltage signals. Excerpt from Figure 3 of the preprint by Wu et al., under CC BY-NC-ND 4.0 license.
Further evidence against prominent local events in dendrites
In contrast to previous work based on two-photon imaging data (Gonzalez et al., 2026; Moore et al., 2025), Wu et al. find that localized “dendrite-only” spikes were surprisingly rare. One potential explanation they bring up is that they record from CA2 dendrites, which are know to be less susceptible to plasticity protocols and therefore might not need as prominent local dendritic signals to instruct plasticity as compared to e.g. CA1 neurons. I find this explanation interesting, but I still have the feeling that the dissociation between earlier findings for local events and this evidence against local events is an interesting point of discussion that still needs to be resolved.
Conclusion
First of all: all of these papers are definitely worth reading! I found in particular the two papers from the Cohen lab to be a very interesting resource with many thought-through analyses and in vivo tests of prior brain slice work. The most surprising finding and the largest discrepancy between this work based on one-photon imaging (Lee et al., 2026; Wu et al., 2026) and prior two-photon prism-based work (Gonzalez et al., 2026; Moore et al., 2025; and other work from the Losonczy lab) is the lack of prominent local dendritic activity observed by the work from the Cohen lab.
It is not clear what could explain this discrepancy. Did the approach to regress out brain motion from voltage imaging signals by Lee et al. suppress true local activity? Did the inserted prism cut off connections, resulting in lower activity levels? (This is actually addressed with a control experiment by Lee et al..) Or may two-photon dendritic recordings be more prone to movement artifacts (despite the huge effort to correct for those) and therefore exhibit prominent apparent local activity? It is known that for typical hippocampal windows, the strata radiatum and lacunosum-moleculare, where the apical dendrites reside, exhibit stronger motion artifacts than soma or basal dendrite; in addition, resolution and SNR are lower for apical recordings, all of this contributing to potentially more motion artifacts in apical dendrites. Other reasons that may explain the difference between findings may be the different methods to induce expression, using single-cell electroporation (which adds a lot of plasmids into a single cell and might lead to overexpression) by Gonzalez et al., and AAV-based expression by Wu et al. and Lee et al. (which may have side-effects due to the injected virus particles).
At this point in time, it seems not yet possible to figure out the technical and biological contributions to motion artifacts or the lack thereof in these studies. So I’m really looking forward to seeing these important experiments replicated and consolidated across other labs!
In any case, check out these papers! I barely scratched the surface, and all of them are definitely worth a read.
References
Alexander, G.M., He, B., Leikvoll, A., Jones, S., Wine, R., Kara, P., Martin, N., Dudek, S.M., 2024. Hippocampal CA2 neurons disproportionately express AAV-delivered genetic cargo. https://doi.org/10.1101/2024.11.27.625768
Beaulieu-Laroche, L., Toloza, E.H.S., Brown, N.J., Harnett, M.T., 2019. Widespread and Highly Correlated Somato-dendritic Activity in Cortical Layer 5 Neurons. Neuron 103, 235-241.e4. https://doi.org/10.1016/j.neuron.2019.05.014
Francioni, V., Padamsey, Z., Rochefort, N.L., 2019. High and asymmetric somato-dendritic coupling of V1 layer 5 neurons independent of visual stimulation and locomotion. eLife 8, e49145. https://doi.org/10.7554/eLife.49145
Francioni, V., Tang, V.D., Toloza, E.H.S., Ding, Z., Brown, N.J., Harnett, M.T., 2026. Vectorized instructive signals in cortical dendrites. Nature. https://doi.org/10.1038/s41586-026-10190-7
Gonzalez, K.C., Noguchi, A., Zakka, G., Yong, H.C., Terada, S., Szoboszlay, M., O’Hare, J., Negrean, A., Geiller, T., Polleux, F., Losonczy, A., 2025. Visually guided in vivo single-cell electroporation for monitoring and manipulating mammalian hippocampal neurons. Nat. Protoc. 20, 1468–1484. https://doi.org/10.1038/s41596-024-01099-4
Gonzalez, K.C., Terada, S., Noguchi, A., Zakka, G.N., O’Toole, C., Bilbao, G., Reynolds, L., Jász, A., Kertész, B., Szadai, Z., Shen, A., St-Pierre, F., Polleux, F., Losonczy, A., Rózsa, B., 2026. Movement-stabilized three-dimensional optical recordings of membrane potential changes and calcium dynamics in hippocampal CA1 dendrites. Neuron S0896627326000048. https://doi.org/10.1016/j.neuron.2026.01.004
Griffiths, V.A., Valera, A.M., Lau, J.Y., Roš, H., Younts, T.J., Marin, B., Baragli, C., Coyle, D., Evans, G.J., Konstantinou, G., Koimtzis, T., Nadella, K.M.N.S., Punde, S.A., Kirkby, P.A., Bianco, I.H., Silver, R.A., 2020. Real-time 3D movement correction for two-photon imaging in behaving animals. Nat. Methods 1–8. https://doi.org/10.1038/s41592-020-0851-7
Larkum, M., 2013. A cellular mechanism for cortical associations: an organizing principle for the cerebral cortex. Trends Neurosci. 36, 141–151. https://doi.org/10.1016/j.tins.2012.11.006
Lee, B.H., Park, P., Wu, X., Wong-Campos, J.D., Xu, J., Xiong, M., Qi, Y., Huang, Y.-C., Itkis, D.G., Plutkis, S.E., Lavis, L.D., Cohen, A.E., 2026. Fast dendritic excitations primarily mediate back-propagation in CA1 pyramidal neurons during behavior. https://doi.org/10.64898/2026.01.03.696606
Major, G., Larkum, M.E., Schiller, J., 2013. Active Properties of Neocortical Pyramidal Neuron Dendrites. Annu. Rev. Neurosci. 36, 1–24. https://doi.org/10.1146/annurev-neuro-062111-150343
Moore, J.J., Rashid, S.K., Bicker, E., Johnson, C.D., Codrington, N., Chklovskii, D.B., Basu, J., 2025. Sub-cellular population imaging tools reveal stable apical dendrites in hippocampal area CA3. Nat. Commun. 16, 1119. https://doi.org/10.1038/s41467-025-56289-9
Poirazi, P., Papoutsi, A., 2020. Illuminating dendritic function with computational models. Nat. Rev. Neurosci. 21, 303–321. https://doi.org/10.1038/s41583-020-0301-7
Redman, W.T., Wolcott, N.S., Montelisciani, L., Luna, G., Marks, T.D., Sit, K.K., Yu, C.-H., Smith, S., Goard, M.J., 2022. Long-term transverse imaging of the hippocampus with glass microperiscopes. eLife 11, e75391. https://doi.org/10.7554/eLife.75391
Rupprecht, P., Carta, S., Hoffmann, A., Echizen, M., Blot, A., Kwan, A.C., Dan, Y., Hofer, S.B., Kitamura, K., Helmchen, F., Friedrich, R.W., 2021. A database and deep learning toolbox for noise-optimized, generalized spike inference from calcium imaging. Nat. Neurosci. 24, 1324–1337. https://doi.org/10.1038/s41593-021-00895-5
Rupprecht, P., Rózsa, M., Fang, X., Svoboda, K., Helmchen, F., 2025. Spike inference from calcium imaging data acquired with GCaMP8 indicators. https://doi.org/10.1101/2025.03.03.641129
Wu, X., Lee, B.H., Park, P., Wong-Campos, J.D., Xu, J., Plutkis, S.E., Lavis, L.D., Cohen, A.E., 2026. A dendrite-resolved, in vivo transfer function from spike patterns to dendritic Ca2+. https://doi.org/10.64898/2026.01.18.700189