This blog post is about algorithms based on deep networks to denoise raw calcium imaging movies. More specifically, I will write about the difficulties to interprete their outputs, and on how to address these limitations in future work. I will also share my own experience with denoising calcium imaging data from astrocytes in mice.
Introduction
Averages from calcium imaging or representative movies in publications often look great. In reality, however, two-photon calcium imaging is often limited primarily by low signal-to-noise ratios. There are many cases where recordings are dominated by shot noise to the extent that almost no structure is visible from a single frame. This can be due to a weakly expressing transgenic line; or on-purpose low expression levels to avoid calcium buffering; or it can be due to the fact that the microscope scans so fast across a large field of view or volume that it only picks up a few photons per neuron.
The advent of new algorithms to denoise calcium imaging movies
So, would it not be great to get rid of the noise in these noise-dominated movies using some magical deep learning algorithm? This seems to be the promise of a whole set of algorithms that were designed to denoise noisy images and recover the true signal using either supervised [1] but recently also self-supervised deep learning [2-3]. Recently, there have also been a few applications of these algorithms for calcium imaging [4-6]. The following tweet by Jérôme Lecoq was the first time I saw the results of such algorithms:
The results look indeed very impressive. The implementations of these algorithms were subsequently published, by Lecoq et al. (DeepInterpolation) [4] and independently with a very similar approach by Li et al. (DeepCAD) [5]. Since then, a few follow-ups were published to compare performance among the two algorithms and also to improve performance of DeepCAD [6], or with ideas to apply a sort of mixture algorithm between supervised and un-supervised to denoise calcium imaging data [7].
Despite the great-looking results, I was a bit skeptical. Not because of a pratical but rather because of a theoretical concern: there is no free lunch. Why should the output suddenly be less noisy than the input? How can the apparent information content increase? Let’s have a closer look at how the algorithm works to address this question.
Taking into account additional information to improve pixel value estimates
Let’s start very simple: Everybody will agree that the measured intensity for a pixel is a good estimate for the true fluorescence or calcium concentration at this location. However, we can refine our estimate very easily using some background information. What kind of background information? For example, if a high intensity value occurs in a region without any labeling, with all other time points of this pixel having almost zero value, we can be rather certain that this outlier is due to a falsely detected photon and not due to a true fluorescence signal. If however all surrounding pixels had high intensity values and the pixel of interest not, we could also correct our estimate of this pixel’s intensity value using (1) our experience about the spatial structures that we want to observe and (2) the information gained from the surround pixels. Therefore, refining our estimate of the pixel’s intensity is simply taking into account a prior what we expect the pixel’s intensity to be.
Methods based on self-supervised deep networks perform more or less such a procedure, and it is in my opinion a very reasonable way to obtain a better estimate for a pixel’s intensity. As a small difference, they only use the surrounding frames (adjacent in time) and not the pixel intensity itself (therefore lacking this Bayesian idea of improving an estimate using prior information). Despite this interesting small difference, it is clear that such denoising will – in principle – work. The network then uses deep learning to gain knowledge about what to expect in a given context; practically speaking, the prior knowledge will be contained in the network’s weights and extracted from a lot of raw data during learning. Using such a procedure, the estimate of the pixel’s intensity value will, probably under most conditions, be better than the raw intensity value.
A side note: Computing the SNR from raw and denoised pixels
From that, it is also clear that neighboring pixels of a denoised movie are correlated since their original values have influenced each other. It is therefore not justified to compare something like a SNR based on single pixels or single frames between raw and denoised data, because in one case (raw data) adjacent data points are truly independent measurements, while in the other (denoised data) they are not. Both DeepInterpolation [4] and DeepCAD [5] used such SNR measures that reflect the visual look and feel but are, in my opinion, not a good quantification of how much signal and how much noise is in the data. But this just as a side note.
Denoising can make plausible point estimates that are however artifacts
However, there is a remaining real problem. Let’s take some time to understand it. Clearly, the estimated intensity value is only a point estimate. So we don’t know anything about the confidence of the network to infer exactly this pixel intensity and not a different intensity value. Deep networks have been often shown to hallucinate familiar patterns when they were unconstrained by input. It is therefore not clear from looking at the results whether the network was very confident about all pixel intensities or whether it just made up something plausible because the input did not constrain the output sufficiently.
To make this rather vague concern a bit more concerete, here is an example of a calcium recording that I performed a few years ago (adult zebrafish forebrain, GCaMP6f). On the left side, you can see the full FOV, on the right side a zoom-in.
In the movie, there is first a version based on raw data, then the same raw data but with a smoothing temporal average, and finally a version denoised using the DeepInterpolation algorithm [4]. To provide optimal conditions, I did not use a default network provided by the authors but retrained it on the same data to which I applied the algorithm afterwards.
First, the apparent denoising is impressive, and it is easy to imagine that an algorithm performing automated source extraction will perform better for the denoised movie as for the raw movie.
When we look more carefully and with more patience, a few odd things pop out. In particular, the neuronal structures seem to “wobble around” a bit. Here is a short extract of a zoom-in into the denoised movie:
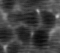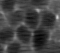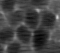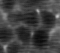
Neurons are densely packed in this region, such that the cytoplasms filled by GCaMP generate an almost hexagonal pattern when you slice through it with the imaging plane. In the excerpt above, there is indeed a sort of hexagonal pattern in each frame. However, the cell boundaries are shifting around from frame to frame. This shifting of boundaries can be particularly well seen for the cell boundary between the right-most neuron and its neighbor to the left. From the perspective of an intelligent human observer, these shifting boundaries are obviously wrong – it is clear that the brain and its neurons do not move.
So, what happened? The network indeed inferred some structural pattern from the noise, but it arrived at different conclusions for different time points. The network made the most likely guess for each timepoint given the (little) information it was provided, but the inconsistency of the morphological pattern shows that the network made up something plausible that however is partially wrong.
Solution (1): Taking into account the overall time-averaged fluorescence
To fix this problem specifically, the network could take into account not only surrounding pixels, but also the overall mean fluorescence (average across all movie frames) in order to make an educated guess about pixel intensity. As human observers, we do this automatically, and that’s why we can spot the artifact to start with. With the information about the overall anatomy, the network would have the same prior as the human observer and would be able to produce outputs that do not include such artifacts.
Solution (2): Taking into account the uncertainty of point estimates
However, the more general problem of the network to fill up uncertain situations with seemingly plausible but sometimes very wrong point estimates still persists. The only difference is that a human observer probably would be unable to identify the generated artifacts.
A real solution to the problem is to properly deal with uncertainties (for reference, here a review of uncertainty in deep networks). This means that the network needs to be able to estimate not only the most likely intensity values for each pixel but also the confidence intervals for each value. With a confidence interval for each pixel value, one could compute the confidence interval for e.g. the time course of a neuron’s calcium ΔF/F averaged across an ROI. The computational problem here is that the error ranges for each pixel do not just add as independent errors, resulting in a standard error of the mean, since the values and confidence intervals for adjacent pixels are dependent on each other. I assume that a straight-forward analytical treatment might be too tricky and some sort of Monte Carlo-based simulation would work better here. This would make it possible to use the denoised movie to derive e.g. a temporal ΔF/F trace of a neuron together with an uncertainty corridor of the estimated trace.
To sum it up, at this point it seems that there is not only a need to develop tools that provide faster and more beautiful denoised images, but even more so procedures to properly deal with uncertainties of estimates that reflect an output that is not enough constrained by the input. Without such tools, analyses based on denoised data must be carefully inspected whether they might be susceptible to such artifacts.
Practical aspects: Using denoising for astrocytic calcium imaging
In a recent preprint [8], I used such methods (DeepInterpolation [4]) to denoise calcium recordings from hippocampal astrocytes. Astrocytes are rather sensitive to laser-induced heating, and I therefore applied low excitation power, resulting in relatively noisy raw recordings. One main goal of the study was to study not only ROIs drawn around somata but also the spatio-temporal calcium dynamics from somatic and distal compartments, ideally with a precision of a single pixel.
To be able to quantify such patterns, it was essential to denoise the raw movie (see Figure 6e and supplemental Figure S8 in [8]). Briefly, it worked really nicely:
It was however crucial to carefully look at both raw and denoised data to understand what was going on, and to consider potential artifacts with respect to downstream analyses. In my case, it helped that the central result of the paper, that calcium signals propagated from distal to proximal compartments under certain conditions, was based on analyses averaged over time (due to the use of correlation functions). Such averaging is likely to undo any harm introduced by small artifacts generated by denoising. In addition, I carefully looked at raw together with denoised data and thought about possible artifacts that might be introduced by denoising.
The second aspect to notice is that the algorithm was rather difficult to use, required a GPU with large memory and still then was very slow. This has improved a bit since then, but the hardware requirements are still high. An alternative algorithm [5] seems to have slightly lower requirements on hardware, and the authors of [5] also developed a modified version of their algorithm that seems to be much faster, at least for inference [6].
Outlook
The development of methods to denoise imaging data is a very interesting field, and I look forward to seeing more work in this direction. Specifically, I hope that the two possible developments mentioned above (taking into account the time-averaged fluorescence and dealing properly with uncertainty) will be properly explored by other groups.
Researchers who apply denoising techniques are themselves often very well aware of potential pitfalls and hallucinations generated by U-Nets or other related techniques. For example, Laine et al. [9] end their review of deep learning-based denoising techniques with this note of caution:
“Therefore, we do not recommend, at this stage, performing intensity-based quantification on denoised images but rather to go back to the raw [images] as much as possible to avoid artefacts.”
With “quantification”, they do not refer to the computation of ΔF/F but rather to studies that quantify e.g. localized protein expression in cells. But should the computation of ΔF/F values have less strict standards?
There are a few cases where potential problems and artifacts are immediately obvious for the application of denoising methods to calcium imaging data. Self-supervised denoising uses the raw data to learn the most likely intensity value given. As a consequence, there will be a tendency to suppress outliers. This is not bad by itself because such outliers are most likely just noise. But there might also be biologically relevant outliers: rare local calcium events on a small branch of a dendritic tree; or unusually shaped calcium events due to intracellularly recruited calcium; or unexpected decoupling of two adjacent neurons that are otherwise strongly coupled by electrical synapses. If the raw SNR is not high enough, the network will take such events as unlikely to be true and discard them in favor of something more normal.
As always, it is the experimenter who is responsible that such concerns are considered. To this end, some basic understanding of the available tools and their limitations is required. Hopefully this blog post helps to make a step into this direction!
References
- Weigert, M., Schmidt, U., Boothe, T., Müller, A., Dibrov, A., Jain, A., Wilhelm, B., Schmidt, D., Broaddus, C., Culley, S. and Rocha-Martins, M. Content-aware image restoration: pushing the limits of fluorescence microscopy. Nature Methods, 15(12). 2018.
- Krull, A., Buchholz, T.O. and Jug, F. Noise2void-learning denoising from single noisy images. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition. 2019.
- Lehtinen, J., Munkberg, J., Hasselgren, J., Laine, S., Karras, T., Aittala, M. and Aila, T. Noise2Noise: Learning image restoration without clean data. arXiv. 2019.
- Lecoq, J., Oliver, M., Siegle, J.H., Orlova, N., Ledochowitsch, P. and Koch, C. Removing independent noise in systems neuroscience data using DeepInterpolation. Nature Methods, 18(11). 2021.
- Li, X., Zhang, G., Wu, J., Zhang, Y., Zhao, Z., Lin, X., Qiao, H., Xie, H., Wang, H., Fang, L. and Dai, Q. Reinforcing neuron extraction and spike inference in calcium imaging using deep self-supervised denoising. Nature Methods, 18(11). 2021.
- Li, X., Li, Y., Zhou, Y., Wu, J., Zhao, Z., Fan, J., Deng, F., Wu, Z., Xiao, G., He, J. and Zhang, Y. Real-time denoising of fluorescence time-lapse imaging enables high-sensitivity observations of biological dynamics beyond the shot-noise limit. bioRxiv. 2022.
- Chaudhary, S., Moon, S. and Lu, H. Fast, Efficient, and Accurate Neuro-Imaging Denoising via Deep Learning. bioRxiv / [Update September 2022: Nature Communications]. 2022.
- Rupprecht, P., Lewis, C., Helmchen, F. Centripetal integration of past events by hippocampal astrocytes. bioRxiv. 2022.
- Laine, R.F., Jacquemet, G. and Krull, A. Imaging in focus: an introduction to denoising bioimages in the era of deep learning. The International Journal of Biochemistry & Cell Biology, 140. 2021.
Hi!
I just want to point out that the link from ref. 6 is wrong.
Still thanks for the great article!
Thanks for noticing! I corrected the wrong link in ref. 6.
Pingback: Why your two-photon images are noisier than you expect | A blog about neurophysiology