Recent Comments
-
Recent Posts
- Subcellular calcium and voltage imaging of pyramidal neurons in mouse hippocampus
- CascadeTorch: a PyTorch version of Cascade for spike inference
- A clearing method for large human FFPE tissue blocks
- Hippocampal neuron and astrocyte responses to noradrenaline and natural arousal
- Annual report of my intuition about the brain (2025, part III)
Archives
- May 2026
- February 2026
- January 2026
- December 2025
- November 2025
- September 2025
- August 2025
- June 2025
- April 2025
- March 2025
- February 2025
- January 2025
- September 2024
- August 2024
- July 2024
- June 2024
- April 2024
- March 2024
- December 2023
- November 2023
- October 2023
- September 2023
- July 2023
- December 2022
- November 2022
- October 2022
- August 2022
- May 2022
- February 2022
- December 2021
- October 2021
- August 2021
- July 2021
- June 2021
- May 2021
- February 2021
- January 2021
- December 2020
- September 2020
- August 2020
- April 2020
- December 2019
- November 2019
- September 2019
- July 2019
- June 2019
- March 2019
- January 2019
- December 2018
- October 2018
- September 2018
- August 2018
- July 2018
- June 2018
- April 2018
- January 2018
- December 2017
- November 2017
- August 2017
- July 2017
- May 2017
- February 2017
- December 2016
- November 2016
- October 2016
- September 2016
- July 2016
- June 2016
- May 2016
- April 2016
- March 2016
- February 2016
- January 2016
- December 2015
- November 2015
- August 2015
- June 2015
- March 2015
- January 2015
- November 2014
- August 2014
- May 2014
- April 2014
- March 2014
- February 2014
Categories
Recent Comments
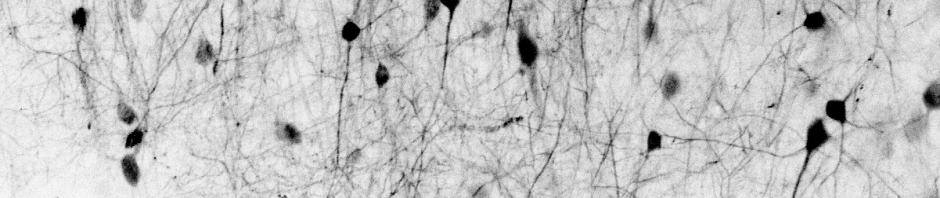
 Reuss AM, Groos D, Cerisoli M, Nordberg L, Frick L, Voigt FF, Vladimirov N, Bethge P, Reimann R, Helmchen F,
Reuss AM, Groos D, Cerisoli M, Nordberg L, Frick L, Voigt FF, Vladimirov N, Bethge P, Reimann R, Helmchen F,  Duss SN, Wilhelm M, Marinescu AM, Zhang R, Helmchen F, Bohacek J†,
Duss SN, Wilhelm M, Marinescu AM, Zhang R, Helmchen F, Bohacek J†, 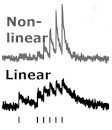
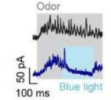 Hu B, Temiz NZ, Chou CN,
Hu B, Temiz NZ, Chou CN, 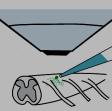
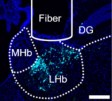 Groos D, Reuss AM*,
Groos D, Reuss AM*, 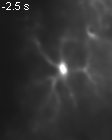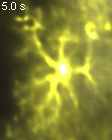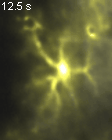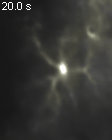
 Masala N, Mittag M, Giovannetti EA, O’Neil DA, Distler F,
Masala N, Mittag M, Giovannetti EA, O’Neil DA, Distler F, 
 Schoenfeld G*, Carta S*,
Schoenfeld G*, Carta S*,  Huang KH,
Huang KH, 
 Berens P, Freeman J, Deneux T, Chenkov N, McColgan T, Speiser A, Macke JH, Turaga S, Mineault P,
Berens P, Freeman J, Deneux T, Chenkov N, McColgan T, Speiser A, Macke JH, Turaga S, Mineault P,  Jacobson GJ,
Jacobson GJ, 

