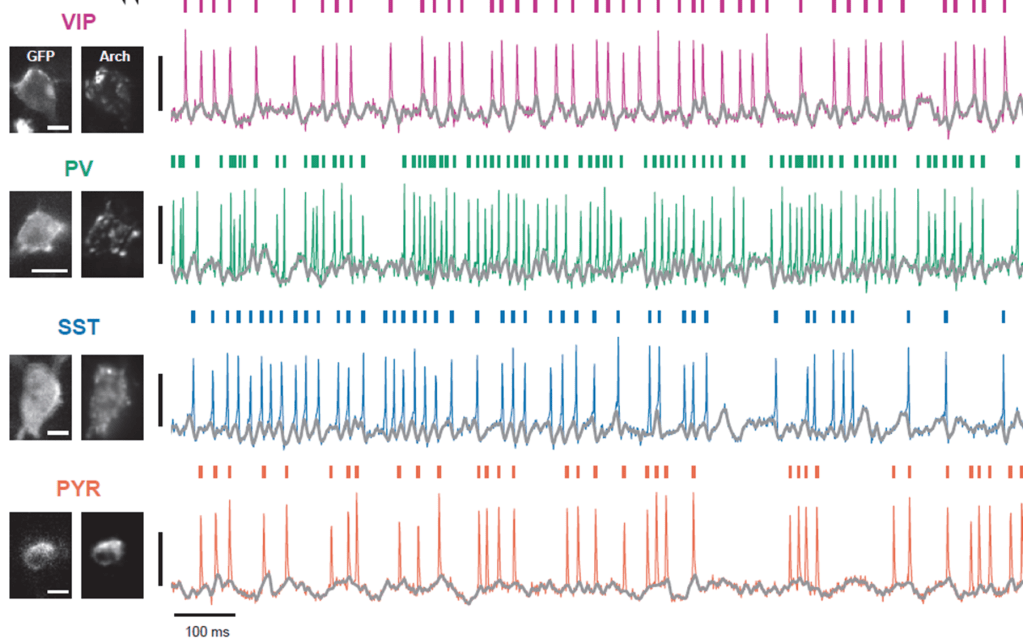
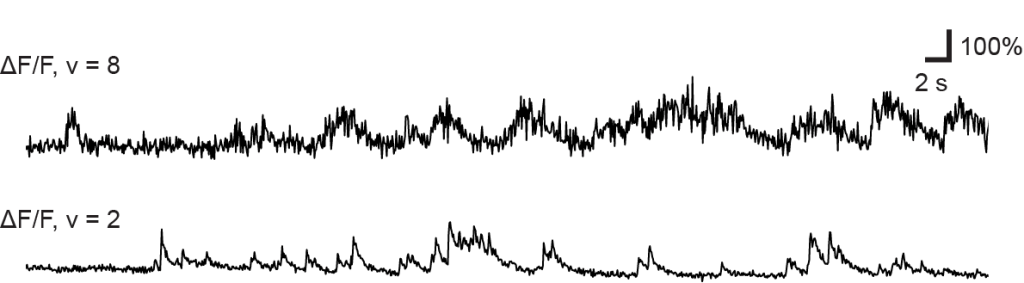
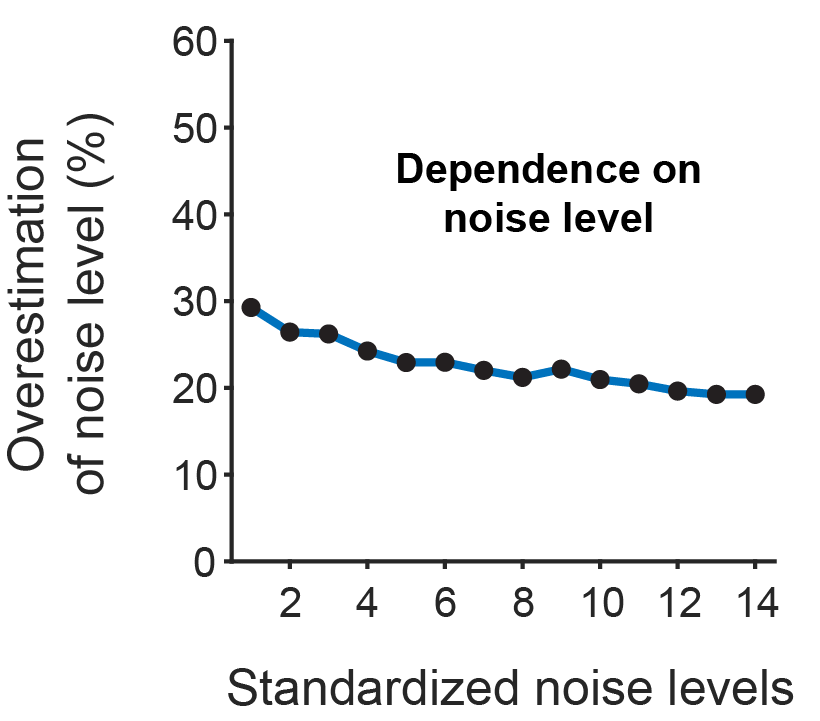
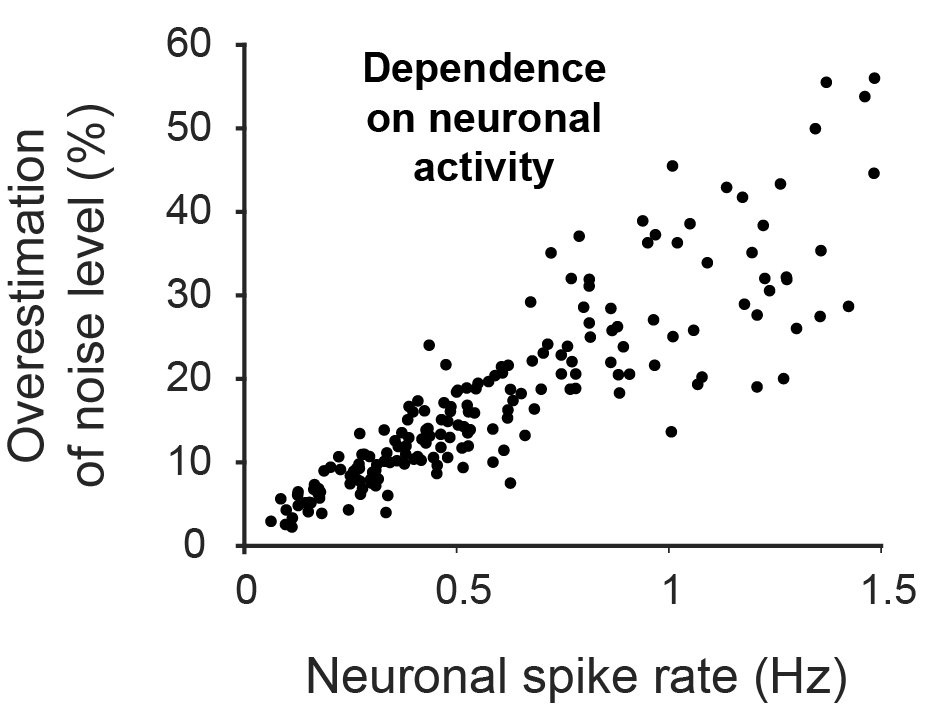
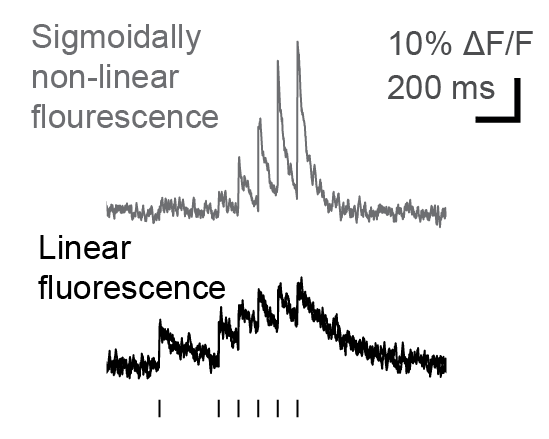
Many researchers, including myself, use calcium imaging to read out the activity of neurons in the living brain. This method relies on the fact an action potential – the “spike” of neuronal activity – opens voltage-gated calcium channels, transiently increasing the calcium concentration in the cell.
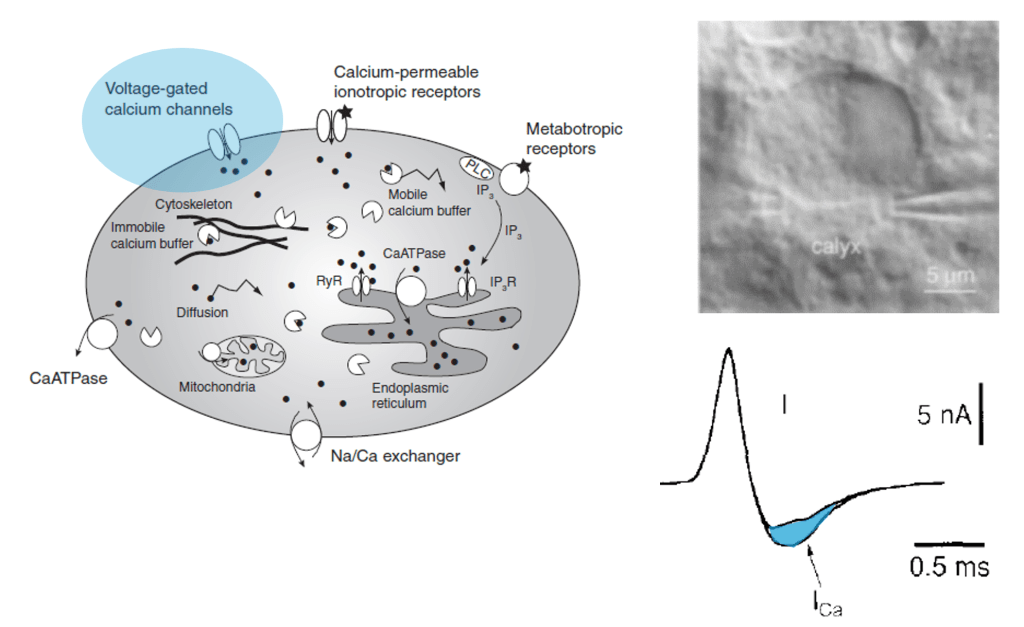
However, calcium isn’t just an indicator of action potentials; it’s a seemingly universal signaling molecule used in most if not all living cells on Earth. In neurons, calcium concentration is also influenced by influx from cell organelles, in particular the endoplasmatic reticulum (ER, a loose web that is weaved through the cytoplasm) and the mitochondria (the “powerhouse” of the cell). These organelles are present not only in the cell body but also in its dendrites and axons. As an expert for calcium imaging (check out my latest work on calcium sensors) with a long-standing interest in dendrites, I thought it would make sense to have a glimpse into the current literature on calcium signals coming from neuronal cell organelles. Here are three papers from the last years that I selected to read more carefully – but I’m happy to receive pointers to additional ones that might be equally relevant!
Calcium signaling at junctions between plasma membrane and endoplasmic reticulum
This paper might be especially interesting for you if you are curious about the role of dendritic spines. In this impressive study, Benedetti et al. (2025) investigate the structural basis of a key player for calcium signaling in neuronal dendrites, the endoplasmic reticulum (ER). They use an amazing array of techniques, from super-resolution lattice light sheet microscopy, volumetric electron microscopy, dual-color calcium imaging in cytosol and ER, extensive pharmacology, to two-photon glutamate uncaging – there is something for almost everybody.
Their main finding is that the ER forms regular junctions with the plasma membrane, spaced by 1 μm, while the rest of the ER is floating in the cytosol like a weave (see figure excerpt below).
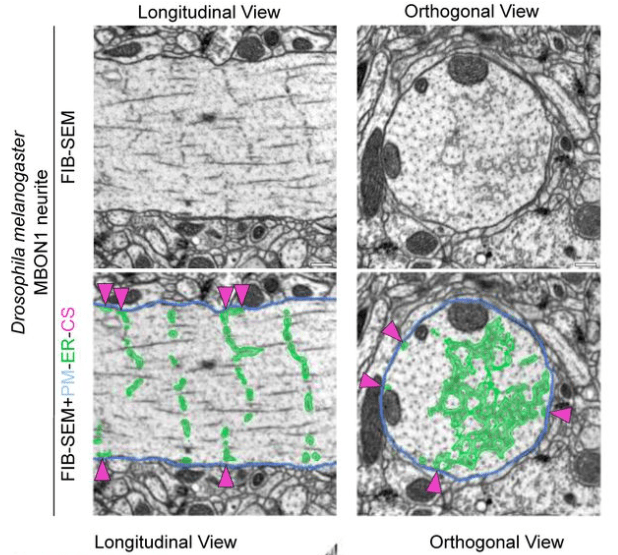
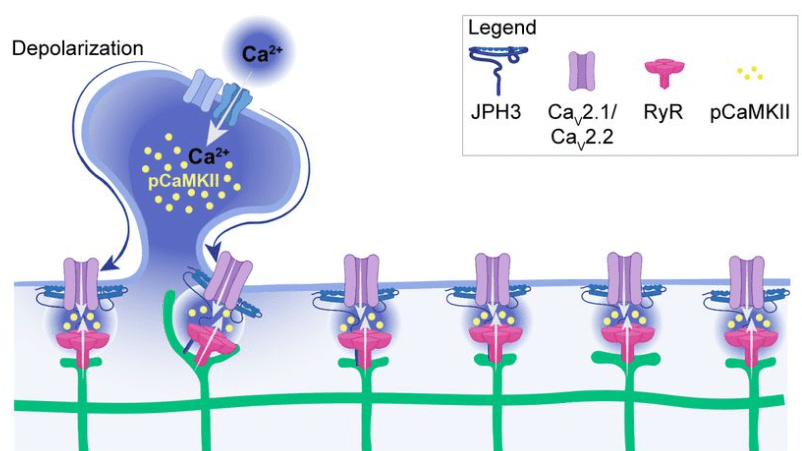
Benedetti et al. show that these junctions are also functionally relevant because they concentrate specific calcium channels and result in local activation of CaMKII. Maybe the most important aspect: The ER can reach into spine heads, and activation of single spines can propagate as calcium signal within the ER up to 20 μm along the dendrite. This observation is important since dendritic spines have often been speculated to be distinct compartments that are, by virtue of the high spine neck resistance, insulated from the neighboring dendritic compartment (that’s still a matter of debate). Here, Benedetti et al. show that the ER can bypass this insulation, not by electrotonic propagation but by calcium signaling.
In summary, a very beautiful paper – where spatially resolved cell biology meets neuroscience.
Cytosolic calcium signals couple only loosely to mitochondrial calcium in vivo
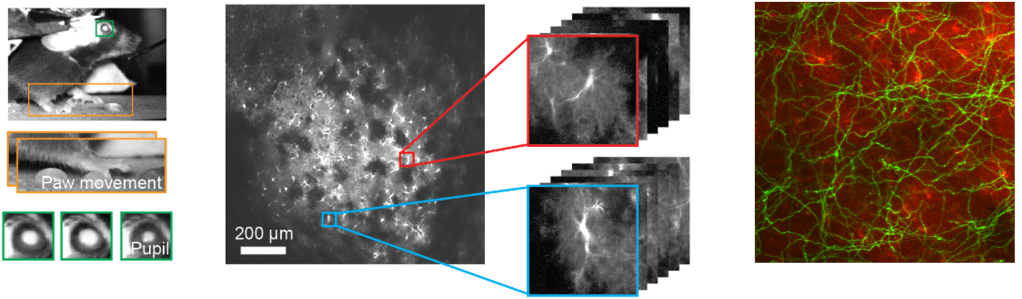
In this interesting work, Lin et al. (2019) study the relationship between cytosolic and mitochondrial calcium signals in the intact mouse brain. To this end, they use dual color-imaging with the green calcium indicator GCaMP expressed in mitochondria and the red-shifted indicator RCaMP in the cytosol.
Previous work from cultured neurons or slices had suggested that there is tight coupling between mitochondrial and cytosolic calcium transients, but these findings were based on artificial stimulation. Here, Lin et al. find a less reliable coupling, with only 3-10% of cytosolic transients being reflected in mitochondrial calcium signaling, with the fraction depending on the processing load of the respective cortical areas (visual or motor cortex). This loose coupling during natural conditions is interesting and also raises the question to which extent artificial stimulation – e.g., the glutamate uncaging experiments performed by Benedetti et al. – relates to physiological stimulation levels.
Lin et al. find a couple of variables to explain which events are coupled and which ones are not. Yet, they conclude that the coupling seems still quite random (“probabilistic”). However, they find that CamKII, a self-phosphorylating kinase also involved in neuronal plasticity, is necessary to enable such coupling.
Another interesting point: Lin et al. find that mitochondrial calcium signals are not limited to small compartments but are synchronized across separate mitochondrial compartments across >15 μm. Since these compartments are only connected by small “nano-tunnels” of the ER, it is likely that calcium signals do not propagate across mitochondrial compartments but are orchestrated by a hidden global factor that could not be clearly observed during this study.
Intracellular calcium release shapes plasticity and place cells in hippocampus
O’Hare et al. (2022) investigate how intracellular calcium release from organelles affects dendritic and calcium signals and plasticity in hippocampal neurons of awake mice.
The key method of this study is single-cell in vivo electroporation, which enables the targeted insertion of plasmids for 1) expression of a calcium indicator, 2) expression of an optogenetic actuator, and 3) activation of a conditional knockout in a single visually targeted pyramidal cell. This method is described in more detail in this recent paper by Gonzalez et al., 2025 from the same lab.
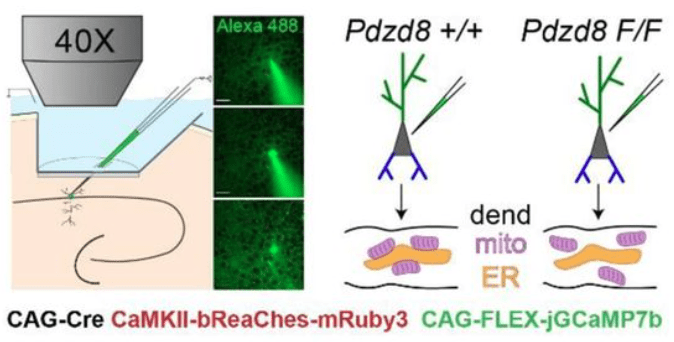
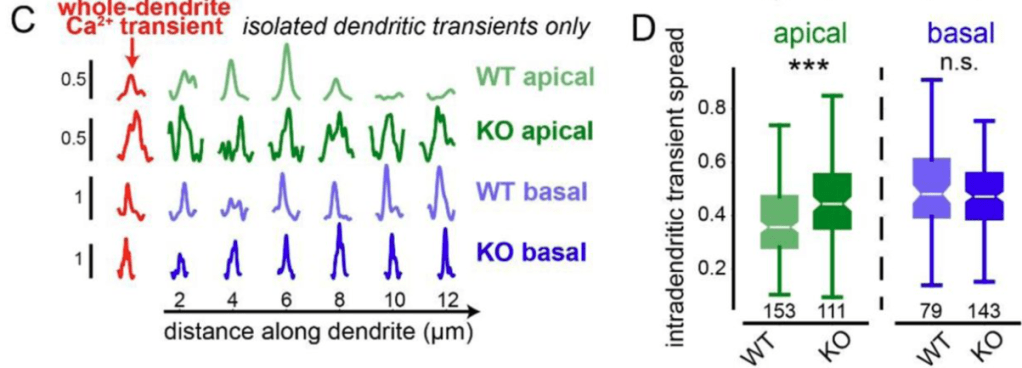
By genetically knocking out a gene responsible for ER-mitochondria coupling (Pdzd8), O’Hare et al. showed that the spatial extent of calcium signaling across the apical dendrites became more widespread (see figure excerpt below). This is a significant finding because extent of calcium signaling coupling between dendritic segments has been a long-standing question that is still not well-understood. This study shows that the functional coupling of intracellular organelles directly influences how calcium signals are compartmentalized within dendrites. In addition, the study demonstrates that this change of intradendritic spread of calcium signals is accompanied by functional plasticity (place cell properties; see the paper for details).
Conclusion
In summary, these papers show an important part of neurons that is often completely forgotten by systems neuroscientists – the cellular biology. A reason why such results are ignored is that they are often obtained in cell cultures or similar artificial systems where the cell organelles and stimulation patterns can be quite different. Lin et al., for example, demonstrate that the tight coupling of mitochondrial to cytosolic calcium is not observed in vivo, underlining the importance of research done in vivo.
A second aspect why there are few such studies is their technical difficulty. Obtaining clean and interpretable signals from two different calcium indicators as done by Lin et al. is very challenging (but see Moret et al., 2025 for the promising development of the chemigenetic sensor HaloCaMP specifically for mitochondria and ER). Similarly, the in vivo electroporation technique by O’Hare et al. is definitely a highly challenging method that I haven’t seen used widely.
Another method where I see a lot of potential for future analyses of cell organelle function in neuronal dendrites is volumetric high-resolution electron microscopy. Existing datasets that are publicly available might contain important information about the interplay of endoplasmic reticulum, mitochondria, the plasma membrane, and the cytosol. Maybe we only need to ask the right question.
.
References
Benedetti, L., Fan, R., Weigel, A.V., Moore, A.S., Houlihan, P.R., Kittisopikul, M., Park, G., Petruncio, A., Hubbard, P.M., Pang, S., Xu, C.S., Hess, H.F., Saalfeld, S., Rangaraju, V., Clapham, D.E., De Camilli, P., Ryan, T.A., Lippincott-Schwartz, J., 2025. Periodic ER-plasma membrane junctions support long-range Ca2+ signal integration in dendrites. Cell 188, 484-500.e22. https://doi.org/10.1016/j.cell.2024.11.029
Gerard, J., Borst, G., Helmchen, F., 1998. Calcium influx during an action potential, in: Methods in Enzymology. Elsevier, pp. 352–371. https://doi.org/10.1016/S0076-6879(98)93023-3
Gonzalez, K.C., Noguchi, A., Zakka, G., Yong, H.C., Terada, S., Szoboszlay, M., O’Hare, J., Negrean, A., Geiller, T., Polleux, F., Losonczy, A., 2025. Visually guided in vivo single-cell electroporation for monitoring and manipulating mammalian hippocampal neurons. Nat Protoc 20, 1468–1484. https://doi.org/10.1038/s41596-024-01099-4
Helmchen, F., 2012. Calcium imaging, in: Brette, R., Destexhe, A. (Eds.), Handbook of Neural Activity Measurement. Cambridge University Press, pp. 362–409. https://doi.org/10.1017/CBO9780511979958.010
Lin, Y., Li, L.-L., Nie, W., Liu, X., Adler, A., Xiao, C., Lu, F., Wang, L., Han, H., Wang, X., Gan, W.-B., Cheng, H., 2019. Brain activity regulates loose coupling between mitochondrial and cytosolic Ca2+ transients. Nat Commun 10, 5277. https://doi.org/10.1038/s41467-019-13142-0
Moret, A., Farrants, H., Fan, R., Zingg, K., Gee, C.E., Oertner, T.G., Rangaraju, V., Schreiter, E.R., De Juan-Sanz, J., 2025. An expanded palette of bright and photostable organellar Ca2+ sensors. https://doi.org/10.1101/2025.01.10.632364
O’Hare, J.K., Gonzalez, K.C., Herrlinger, S.A., Hirabayashi, Y., Hewitt, V.L., Blockus, H., Szoboszlay, M., Rolotti, S.V., Geiller, T.C., Negrean, A., Chelur, V., Polleux, F., Losonczy, A., 2022. Compartment-specific tuning of dendritic feature selectivity by intracellular Ca2+ release. Science 375, eabm1670. https://doi.org/10.1126/science.abm1670