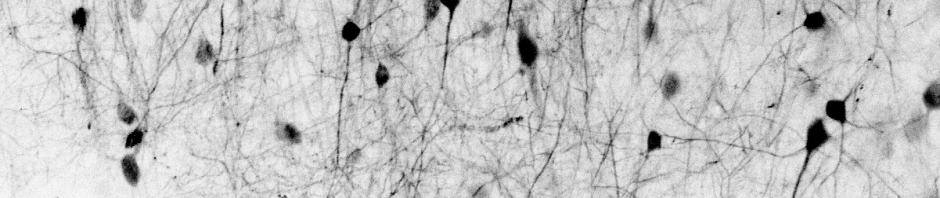
Some time ago, Stephan Gerhard and I have used a convolutional neural network (CNN) to detect neuronal spikes from calcium imaging data. (I have mentioned this before, here, here, and on Github.)
This method is covered by the spikefinder paper that was recently published (Berens et al., 2018), based on a competition that featured ground truth for a training and a test dataset. This was a great and useful competition. But there are some important caveats (which are mentioned in the discussion of the main paper). Here, I will discuss one of the caveats.
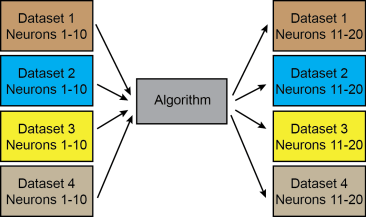
For the competition, the test set consisted of five datasets; neurons not included in the test set, but from the same original datasets with similar conditions and signal-to-noise etc. had been included in the training datasets. Therefore, the competition does not allow to understand how well the networks would generalize to neurons from datasets that have not been used for training. In theory, there would be one algorithm to learn everything and to predict everything (case 1):
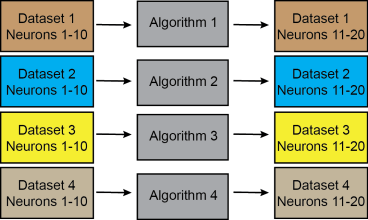
But of course it was possible to e.g. train one method for each dataset separately (case 2), without violating the conditions of the competition (the left column symbolizes the training datasets, the right column the test datasets, colors indicate the same original datasets):
And of course there is a smooth transition between the first and the second possibility, such that it is not easy to judge how closely an algorithm has been fine-tuned for the respective datasets.
Fine-tuning would allow to perform very well in the competition with an algorithm similar to case 2, but there would be no means to easily generalize the method to unseen datasets – and that’s not really desirable.
It is obvious why the competition necessarily had this ‘design flaw’: To circumvent this problem, the training set would have to consist of maybe 15-20 datasets of a couple of neurons each, and the test dataset of ~10 independent datasets (this would be my own rough estimate). Currently, nobody has this amount of ground truth data.
However, I’m interested in applying the prediction method that we developed to new calcium imaging, so I wanted to know how well the network generalizes. Partially, this has already been investigated by Pachitariu et al. (bioRxiv, 2017), but this study did not consider the test datasets directly (which were not public at this point in time).
So I basically trained the CNN algorithm (this one) on all datasets, except on the one I wanted to use as test dataset. Something like this (case 3):
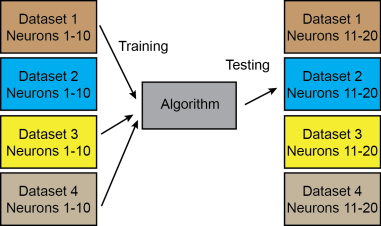
Then I compared the result with the predictions from the normal training (‘case 1’) for the test data. Here are the results:
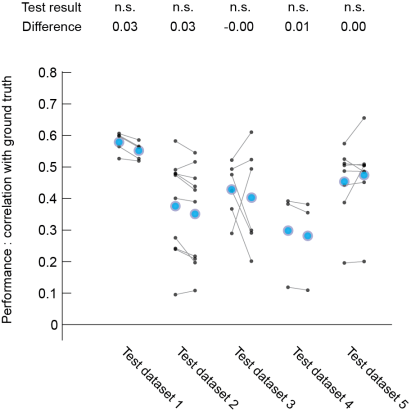
Each black dot corresponds to the prediction performance for one neuron (the greater the value, the better). The left-hand datapoints for each of the five datasets are results from a network that has been trained on all datasets (case 1), whereas right-hand datapoints are results from a network that has not been trained with other neurons from the same original dataset (case 3). Blue dots indicate the mean across neurons for each dataset. The difference indicated at the top is the pseudomedian difference (‘case 1’ minus ‘case 3’), the test used is a non-parametric one.
Judging from this plot, I would say that the predictions are maybe slightly worse, but not much for ‘case 3’ compared to ‘case 1’ (no statistical differences found, but I think there is a tendency). This is pretty reassuring with respect to the power of the algorithm/network to generalize to unseen datasets – it works.
Update September 2020: A much more rigorous and complete analysis of this problem, now on bioRxiv.