Recently, I mentioned a public competition for spike detection – spikefinder.codeneuro.org. I decided to spend a day two days and have a closer look at the datasets, especially the training datasets that provide both simultaneously recorded calcium and spike trains for single neurons. In the following paragraphs, I will try to convey my impression of the dataset, and I will show some home-brewed and crude, but nicely working attempts to infer spiking probabilities from calcium trains. [Update: Together with Stephan Gerhard, I’ve designed a better algorithm based on CNNs to infer spiking probabilities, described on this blog and on Github.]
The training dataset consists of 10 separate datasets, recorded in different brain regions and using different calcium indicators (both genetically encoded and synthetic ones), each dataset with 17±10 neurons, and each neuron recorded for several hundred seconds.
To check the quality of the recordings, I calculated the cross-correlation function between spike train and calcium trace to get the shape of the typical calcium event that follows a spike (it is kind of similar to a PSTH, but takes also cases into consideration where two spikes occur at the same time). I have plotted those correlation functions for the different datasets, one cross-correlation shape for each neurons (colormap below, left). Then I convolved the resulting shape with the ground truth spike train.
If calcium trace and spike train were consistent, the outcome of the convolution would be highly correlated with the measured calcium signal. This is indeed the case for some datasets (e.g. dataset 6; below, right). For others, some neurons show consistent behavior, whereas others don’t, indicating bad recording quality either of the calcium trace or the spike train (e.g. the low correlation data points in datasets 2, 4, 7 and 9).
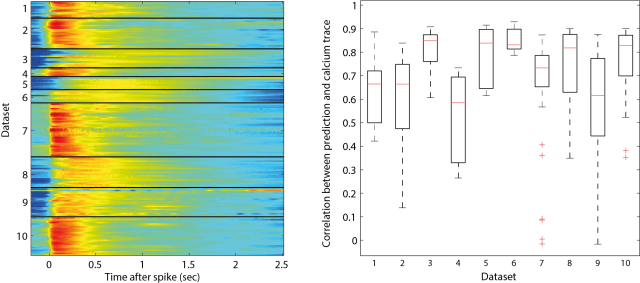
In my opinion, those bad recordings should have been discarded, because there is probably no algorithm in the world that can use them to infer the underlying spike times. From looking at the data, I got the impression that it does not really make sense to try to deconvolve low-quality calcium imaging recordings as they are sometimes produced by large-scale calcium imaging.
But now I wanted to know: How difficult it is to infer the spike rates? I basically started with the raw calcium trace and tried several basic operations to find something that manages to come close to the ground truth. In the end, I used a form of derivative, by subtracting the calcium signal of a slightly later (delayed) timepoint from the original signal. I will link the detailed code below. I was surprised how little parameter tuning was required to get quite decent results for a given dataset. Here is the core code:
% calculate the difference/derivative prediction = calcium_trace - circshift(calcium_trace,delay); % re-align the prediction in time prediction = circshift(prediction,-round(delay/2)); % simple thresholding cut-off prediction( prediction < 0 ) = 0;
Let me show you a typical example of how the result looks like (arbitrary scaling):

Sometimes it looks better, sometimes worse, depending on data quality.
The only difficulty that I encountered was the choice of a single parameter, the delay of the subtracted calcium timetrace. I realized that the optimal delay was different for different datasets, probably due to different calcium indicator. Presumably, this reflects the fact that for instance calcium traces of synthetic dyes like OGB-1 (bright baseline, low dF/F) look very different from GCaMP6f traces (dim baseline, very high dF/F). Those properties can be disentangled e.g. by calculating the kurtosis of the calcium time trace.
Although this is based on only 10 datasets, and although I do not really know the reason why, the optimal delay in my algorithm seemed to depend on the kurtosis of the recording in a very simple way that could be fitted by a simple function (e.g. a double-exponential, or simply a smooth spline):

In the end, this algorithm is barely longer than 20 lines in Matlab (loading of the dataset included), in the simplicity spirit of the algorithm suggested by Dario Ringach. Here’s the full algorithm on Github. I will also submit the results to the Spikefinder competition in order to see how good this simple algorithm is compared to more difficult ones that are based on more complex models or analysis tools.
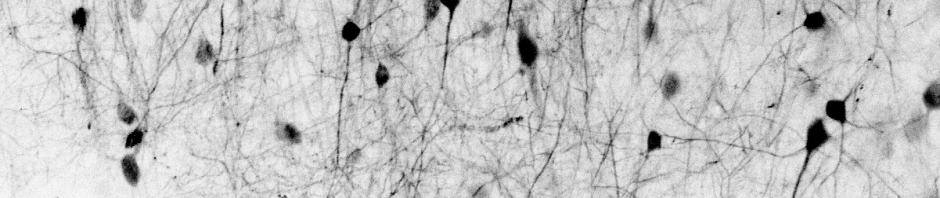
Yes, there are a few crappy recordings…
Pingback: A convolutional network to deconvolve calcium traces, living in an embedding space of statistical properties | A blog about neurophysiology