As a short modeling session for an electrophysiology course at the University of Zurich, I made a tutorial for students to play around with the Hodgkin-Huxley equations in a Colab Notebook / Python, which does not require them to install Python. You’ll find the code online on a small Github repository: https://github.com/PTRRupprecht/Hodgkin-Huxley-CC-VC
Using the code, the students can not only play around with the Hodgkin-Huxley equations, but also replicate in silico the experiments they have done when patching cells in slices (including voltage clamp experiment).
It is really rewarding to be able to reproduce current clamp (recording the membrane potential) and voltage clamp experiments (recording the currents while clamping the voltage to a constant value), because this also allows to replicate computationally the experiments and plots generated experimentally by Hodgkin and Huxley.
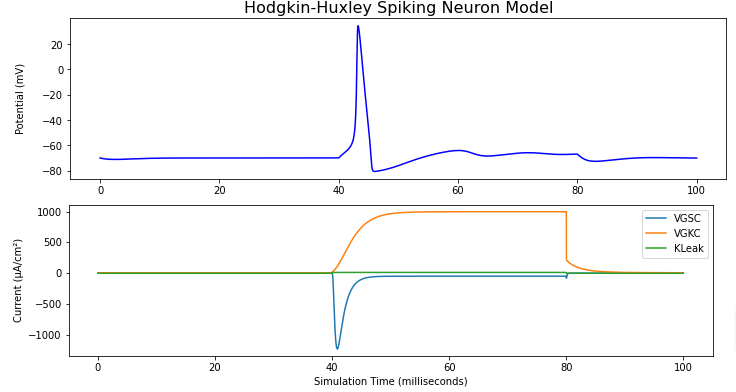
Below, you see a part of the code, the result of a simulation of a Hodgkin-Huxley model. The top configuration was run in current clamp, with a current pulse injected between 40 and 80 ms, which triggered a single action potential. The lower configuration was run in voltage clamp, with the holding potential stepping from -70 mV to -30 mV between 40 and 80 ms. You can clearly see the active conductanves (deactivating sodium conductance in blue and non-deactivating potassium conductance in orange):