How does the noisiness of the recorded calcium data affect the performance of spiking-inferring deconvolution algorithms? I cannot offer a rigorous treatment of this question (Update August 2020: Now I have treated this question rigorously.) , but some intuitive examples. The short answer: If a calcium transient is not visible at all in the calcium data, the deconvolution will miss the transient as well. It seems that if the signal-to-noise drops below 0.5-0.7, the deconvolution quickly degrades.
To make this a bit more quantitative, I used an algorithm based on convolutional networks (developed by Stephan Gerhard and myself; you can find it on Github, and it’s described here) and a small part of the Allen Brain Observatory dataset.
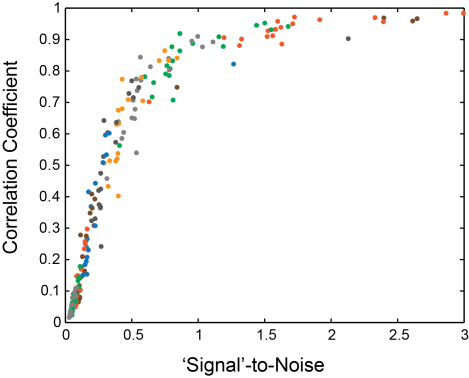
I assumed that the standard deviation of the raw calcium traces measures ‘Signal’ (a reasonable approximation), and I took the standard deviation of the Gaussian noise that I added on top as ‘Noise’. Then I deconvolved both noisified and unchanged calcium traces and computed the correlation of the spiking traces of calcium+noise vs. calcium alone. If the correlation (y-axis) is high, the performance of the algorithm is not much affected by the noise. The curve is dropping steeply at a SNR of 0.5-0.7.
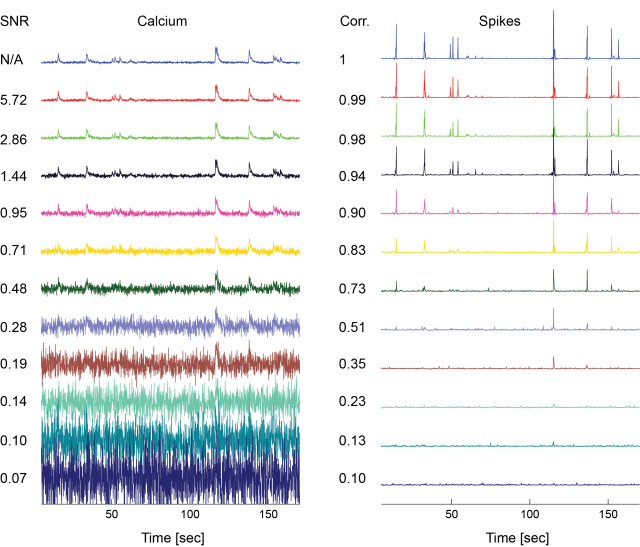
To get some intuition, let’s give some examples, left the calcium trace plus Gaussian noise, right the deconvolved spiking probabilities (numbers to the left indicate SNR and correlation to ground truth, respectively):
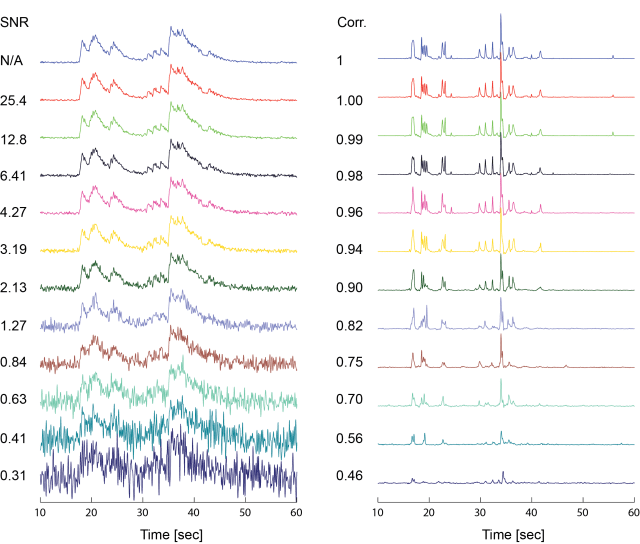
The next example was perturbed with the same absolute amount of noise, but due to the larger signal, the spike inference remained largely unaffected for all but the highest noise levels.
The obvious thing to note is the following: When transients are no longer visible in the calcium trace, they disappear in the deconvolved traces as well. I’d also like to note that both calcium timeseries from the examples above are from the same mouse, the same recording, and even the same plane, but the SNR of the recordings is a lot different. Therefore, lumping together neurons of the same recording, but of different recording quality combines different levels of detected detail. An alternative way would be to set a SNR threshold for the neurons to be included – depending on the precision required from the respective analysis.
Pingback: How well do CNNs for spike detection generalize to unseen datasets? | A blog about neurophysiology
Hi Peter. Thank you so much for your excellent blog! I wonder what is the standard way to calculate the SNR of the calcium signal.
Good question! I have written a blog post about a question more recently: https://gcamp6f.com/2025/04/25/accuaretly-computing-noise-levels-for-calcium-imaging-data/
In this blog post, I show how to calculate the noise level, which is already a good metric. If you want to calculate the signal-to-noise level, you can take this noise level metric, but you also have to define what your signal is. Is it the peak amplitude of your calcium transients? Or the minimal detectable calcium transient? This will depend on the question that you ask scientifically.